Protein Profile #1: RNA polymerase II
Much like the variety of outfits we wear to follow the latest fashion trend, cells mediate their own styles through changing their gene expression patterns to fit with their surrounding environment. Gene expression is an important daily event for a cell which results in the production of proteins but also non-coding products such as functional RNAs. Transcription is the vital first step of gene expression that copies a segment of DNA into RNA catalysed by the enzyme RNA polymerase (RNAP). We, like the rest of eukaryotes have three different types of RNAPs (I, II and III) that synthesise different types of RNA.
RNAPII is the DNA dependent RNA polymerase whose primarily responsible for synthesising messenger RNA (mRNA) that serves as the code for making proteins. Recent work has highlighted some interesting physicochemical features about RNAPII, including the ability to form liquid-like droplets.
Structure
RNAPII is composed of several subunits, 12 to be exact in humans, that interact to form the quaternary structure. The largest of the subunits is RPB1, which forms the DNA binding domain along with several other subunits (Figure1).

Figure 1 RNAPII
RPB1 contains a very important domain, the Carboxy-terminal domain (CTD) so named because it is at the C-terminal of the protein (Figure 2). The CTD consists of several heptad repeats of the following amino acid sequence; Tyr, Ser, Pro, Thr, Ser, Pro, Ser (YSPTSPS). Each residue in the repeat is typically referred to by the numerical position in the repeat, i.e Thr, would by Thr4. These repeats stretch out 52 times in humans and 26 in budding yeast, however the pattern of amino acids becomes less conserved the further out you go. The CTD is not required for the catalytic functions of transcription, but plays a very important role in the regulation.

Figure 2 The anatomy of a protein
A landing pad of interactions
Unlike the typical intricately folded nature of proteins, the CTD is a largely unstructured polypeptide chain, allowing the domain to act as a ‘landing pad’ for the binding of various nuclear factors. These include transcription factors, chromatin modifiers and RNA processing enzymes. Each of the residues in the heptad repeat can be dynamically modified through phosphorylation, proline isomerisation and O-GlcNAcylation, which alters the interactions that can form with these proteins.
A CTD code
Taking into account the number of repeats and the potential for each residue to be phosphorylated you can quite easily imagine multiple versions of the CTD with different binding proteins. In reality, the code is not as astronomical in combination of as may be expected, since the majority of repeats at any time are unphosphorylated or only singly phosphoylated. The CTD code is still nonetheless very important for transcriptional regulation.
A tale of two serines
Although each residue can be modified, most research has primarily focused on Ser2 and Ser5 since the stages they are phosphorylated correspond nicely with the stages of transcription; initiation, elongation and termination.
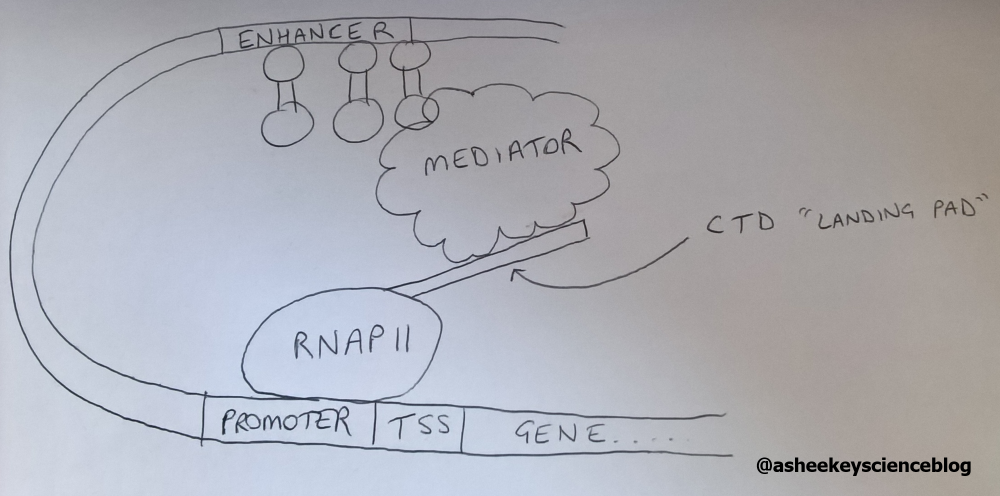
The CTD starts of initially unphosphorylated. This is to allow binding of the Mediator complex, a collection of proteins that also aids transcriptional activation ‘mediating’ interactions between enhancers (binding sites of transcription factors) and promoters (Figure 3).
Figure 3 The mediator bound to the CTD ‘mediating’ enhancer-promoter interactions
Ser5 phosphorylation (Ser5P) by a general transcription factor (GTF), TFIIH, aids promoter escape, hence Ser5P levels are high at the promoter and transcriptional start site (TSS) of a gene. This modification aids in the recruitment of the mRNA capping complex which is a crucial step for later mRNA processing. However, RNAPII is not quite free yet to transcribe the complete gene as a key regulatory event is promoter-proximal pausing which occurs 20-100bp downstream of the TSS. This acts as a transition point between early and productive elongation.
To get passed this step a number of residues on several proteins need phosphorylating, including Ser2. This is achieved by P-Tefb (the positive transcription elongation factor) which through phosphorylation causes proteins preventing elongation to leave. There is a drop in the levels of both Ser2P and Ser5P by the end of the gene hinting that this may aid transcriptional termination. This changing pattern of phosphorylation involving Ser2 and Ser5 is conserved across many species.
For more information of the dynamic phosphorylation pattern of the other residues see Figure 2 of Ref(1).
A transcriptional bubble?
A recent phenomenon discovered in cells is the formation of liquid-like droplets that act as specialised compartments of protein complexes and other components. This has been termed liquid-liquid phase separation and allows cells to separate and localise biochemical reactions from the rest of the cytoplasm.
How these droplets form is still not particularly clear, but one reoccurring feature is the presence of low complexity and intrinsically disordered protein domains. Not only does the CTD contain low complexity domains, but so does two transcription related proteins, FUS and TAF15 which both enable phase separation.
Various CTD repeat lengths where tested in vitro to see if they could be incorporated into the hydrogel formed by TAF15. Interestingly, the minimum CTD-length needed was 8-10 repeats which corresponds to the minimum length required for viability in yeast. This may suggest that the formation of a ‘transcriptional droplet’ is another mechanism necessary for activation of gene transcription. The CTD had to be unphosphorylated to be incorporated matching with the CTD state at the start of transcriptional initiation.
It is unexplored if the CTD is involved in liquid phase separation after transcriptional initiation, but this will be an exciting area of further research.
Further Reading:
(1) K.M.Harlem. The code and beyond: transcription regulation by the RNA polymerase II carboxy-terminal domain Nat Rev Mol Cell Biol (2017) 4, 263-273
3 thoughts on “Get transcribed in style: The CTD that everyone’s talking about”